.
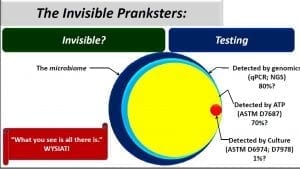
I’m starting this post with an illustration from one of my recent presentations (click on the image to enlarge it). The quote is from Daniel Kahneman’s book: Thinking, Fast and Slow. It reminds us of how often our perceptions are much more limited than we realize. Let’s turn to the circles to the right of the quote.
Now let’s look at the four circles to the left. The large blue circle represents the microbiome – all of the microbes present in a particular environment. Although this post’s focus is fuel systems, some readers might have come across recent reports about the human microbiome. Same concept.
Now let’s look at the tools available for understanding the microbiome. Right now, the most promising tool is Next Generation Sequencing – NGS. This approach is based on extracting all of the genetic material (deoxyribonucleic acid – DNA) from a sample, using enzymes and heating/cooling cycles to multiply the concentration of each type of DNA, separating each type of DNA and then comparing it against a giant DNA library. NGS probably detects >80% of the total population (in the words of Donald Rumsfeld: we have unknown unknowns), but it is fairly expensive and is therefore used as a research tool. There are several What’s New posts about ATP testing. I estimate that ASTM D7687 detects approximately 70% of all microbes in a fuel system sample, however, because the ATP in all cells is the same molecule, ATP testing only provides total bioload data. ATP results won’t tell you what kind of microbes are present (but see my October 2016 post: PROTOCOL FOR DIFFERENTIATING BETWEEN BACTERIAL AND FUNGAL ATP NOW PART OF ASTM E2694)
Contrast NGS and ATP detection with the most commonly used test; culture testing. The story the little red dot (I used 1% because a smaller dot wouldn’t be visible; my real estimate is that culture testing detects between 0.01 and 0.1% of the microbes in a fuel or fuel-associated water sample) tells us is that if you rely on culture testing, you have a >99% chance of failing to detect microbial contamination that might be present in your sample. I’ll explain this more in my next post. Stay tuned. Also, if you have specific questions that you’d like me to address in a future post, please drop me a line at fredp@biodeterioration-control.com